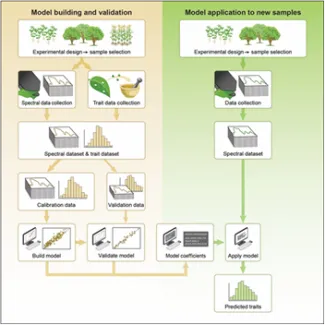
Best practices, a computer package, and hands-on tutorials to guide the application of a powerful technique for prediction of plant traits from hyperspectral data.
The estimation of leaf traits from hyperspectral reflectance data enables rapid, high-throughput, non-destructive characterization of leaf function and plant phenotyping with applications in ecosystem characterization and monitoring. The use of partial least squares regression (PLSR) modeling has been demonstrated to be a powerful tool for leaf trait prediction using hyperspectral data, with recent work demonstrating successful prediction of numerous structural, biochemical, and physiological traits across a wide range of different plant species. However, a lack of consensus on the approach and reporting of results has presented challenges to the wider application of this technique; model results may be difficult to interpret, and the application of published models to new data sets nearly impossible. Scientists with the NGEE Arctic project and others provide a detailed description of use of PLSR to predict leaf traits from hyperspectral data with recommendations for best practice to use across all steps of the process, from experimental design, data collection, PLSR model building, model application and result reporting. These authors advocate for a more standardized workflow to the use of PLSR for prediction of plant traits, with the goal of improving model robustness and enabling comparison of results across research groups, datasets, projects, and biomes. The paper is accompanied by detailed example R-based tutorials to assist users to understand the PLSR model development process and apply the technique and best-practices to their own data.
Citation: Burnett, A. C., J. Anderson, K. J. Davidson, K. S. Ely, J. Lamour, Q. Li, B. D. Morrison, D. Yang, A. Rogers, and S. P. Serbin. 2021. “A best-practice guide to predicting plant traits from leaf-level hyperspectral data using partial least square regression.” Journal of Experimental Botany erab295. https://doi.org/10.1093/jxb/erab295.